Siemer Lab: Biomolecular Solid-State NMR at USC

| Email: | asiemer@usc.edu |
| Office Location: | ZNI room 119F |
| Office Phone: | (323) 442 2720 |
| Lab Location: | ZNI 113 |
| Lab Phone: | (323) 442 2719 |
| Website: | siemerlab.usc.edu |
| Ph.D. Programs: | PIBBS |
The aim of my laboratory is to create a better understanding of the pathological and functional aspects of protein disorder and aggregation. In particular, we are interested in the differences between functional and pathological amyloid fibrils and the importance of intrinsically disordered protein domains (i.e. domains that lack stable tertiary or secondary structure) for the formation of amyloid.
Amyloid formation is important for many neurodegenerative diseases such as Alzheimer's, Parkinson's, and Huntington's disease. The aggregation of monomeric proteins into a variety of oligomeric and fibrillar (amyloid) states can lead to cell death and, therefore, disease. However, amyloid fibrils can also have non-pathological, positive functions such as cell signaling or scaffolding functions. One of the most important goals of the lab is to understand what distinguishes functional and pathological amyloid fibrils. Understanding the difference between these types of amyloid could lead to the development of new therapeutics and will deepen our understanding of the role of amyloid in biology.

Model of an amyloid fibril
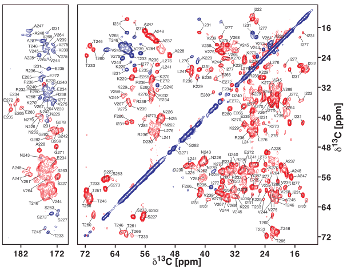
Solid-state NMR spectrum of an amyloid fibril
second area of research in my laboratory is protein disorder as prerequisite for amyloid formation. Intrinsic disorder or partial unfolding is necessary for a protein to aggregate from its soluble state into an amyloid fibril. Furthermore, many proteins can be forced to fold into an amyloid by partial denaturation in vitro. The structural aspects of the unfolded state is, therefore, important for the understanding of amyloid formation and may lead to new strategies to prevent the formation of amyloid in neurodegenerative diseases. However, partially unfolded, intrinsically disordered proteins also carry important functional aspects, especially in eukaryotic organisms, where up to 40% of all proteins contain disordered regions. Our second goal is to understand how aggregation is prevented in most disordered proteins and why it occurs in some disease-related cases.
To address these questions, my lab will use primarily solid-state nuclear magnetic resonance (NMR) in conjunction with other biochemical methods. Solid-state NMR is an emerging technique that enables us to investigate protein structures that are otherwise inaccessible to atomic-resolution structural methods such as X-ray crystallography and liquid-state NMR, thereby permitting us to investigate systems such as protein fibrils with atomic resolution.

